Wyatt Yue
Structural biologist


I hold a chair at the Newcastle University Biosciences Institute as Professor of Structural Biology. I lead a team of postdoc and graduate researchers in the Yue lab, integrated into the Newcastle Structural Biology Laboratory. My research focuses on a moleulcar understanding towards the cause and therapy for rare metabolic diseases within the research theme of Discovery of Medicines, and aligns with the Newcastle Centre for Rare Diseases.
wyatt [dot] yue [at] newcastle [dot] ac [dot] uk
Biosciences Institute
Newcastle University
Catherine Cookson Building M3.008, Framlington Place, Newcastle NE2 4HH
Dynamic inter-domain transformations mediate the allosteric regulation of human 5, 10-methylenetetrahydrofolate reductase
2024
Blomgren LKM, Huber M, Mackinnon SR, Bürer C, Baslé A, Yue WW, Froese DS, McCorvie TJ
Nat Commun
15,
3248
Architecture and regulation of filamentous human cystathionine beta-synthase
2024
McCorvie TJ, Adamoski D, Machado RAC, Tang J, Bailey HJ, Ferreira DSM, Strain-Damerell C, Baslé A, Ambrosio ALB, Dias SMG, Yue WW
Nat Commun
15,
2931
Molecular basis for the regulation of human glycogen synthase by phosphorylation and glucose-6-phosphate
2022
McCorvie TJ, Loria PM, Tu M, Han S, Shrestha L, Froese DS, Ferreira IM, Berg AP, Yue WW
Nat Struct Mol Biol
29,
628-638
Fragment Screening Reveals Starting Points for Rational Design of Galactokinase 1 Inhibitors to Treat Classic Galactosemia
2021
Mackinnon SR, Krojer T, Foster WR, Diaz-Saez L, Tang M, Huber KVM, von Delft F, Lai K, Brennan PE, Arruda Bezerra G, Yue WW
ACS Chem Biol
16,
586-595
Human aminolevulinate synthase structure reveals a eukaryotic-specific autoinhibitory loop regulating substrate binding and product release
2020
Bailey HJ, Bezerra GA, Marcero JR, Padhi S, Foster WR, Rembeza E, Roy A, Bishop DF, Desnick RJ, Bulusu G, Dailey HA Jr, Yue WW
Nat Commun
11(1),
2813
Structure of the human frataxin-bound iron-sulfur cluster assembly complex provides insight into its activation mechanism
2019
Fox NG, Yu X, Feng X, Bailey HJ, Martelli A, Nabhan JF, Strain-Damerell C, Bulawa C, Yue WW, Han S
Nat Commun
10(1),
2210
Palladium-mediated enzyme activation suggests multiphase initiation of glycogenesis
2018
Bilyard MK, Bailey HJ, Raich L, Gafitescu MA, Machida T, Iglésias-Fernández J, Lee SS, Spicer CD, Rovira C, Yue WW, Davis BG
Nature
563,
235-240
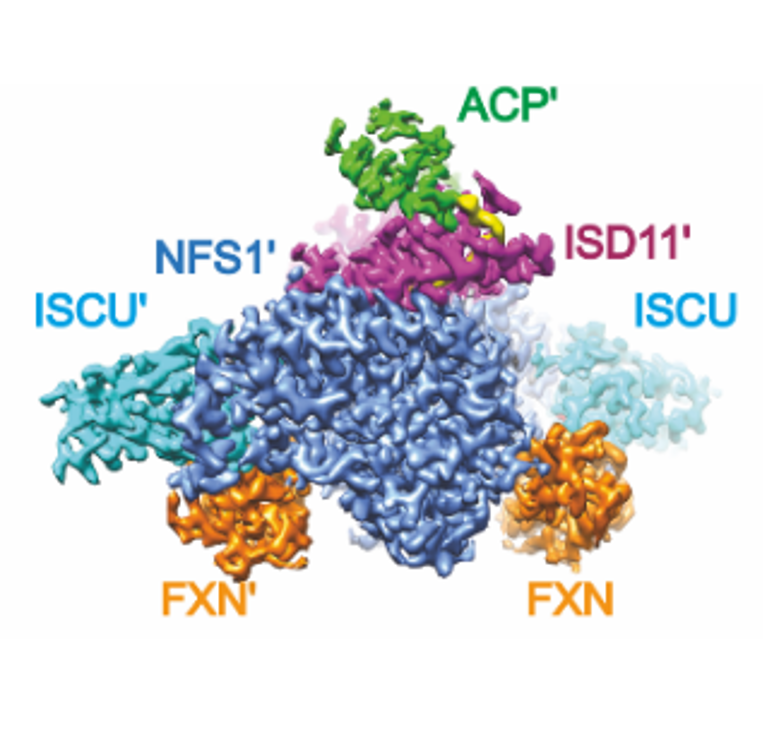
ENZYME ASSEMBLIES & COMPLEXES
Many metabolic enzymes performing essential cellular roles function as homo-oligomeric assemblies, or in concert with other partner proteins in the cell. Therefore studying enzymes in the context of their higher order complexes is key to understanding functions. We adopt novel approaches in recombinant co-expression and endogenous isolation of metabolic complexes, for structural, biochemical and biophysical characterisation.

INTEGRATIVE STRUCTURAL BIOLOGY
We combine x-ray crystallography, cryo-electron microscopy and complementary biophysical methods (including differential scanning fluorimetry, surface plasmon resonance, Michaelis-Mention kinetics) to study enzyme structures, dynamics and interactions. We have to date determined > 200 structures, all deposited in the Protein Data Bank. Our goal is to provide insight into functional and disease-causing mechanisms.
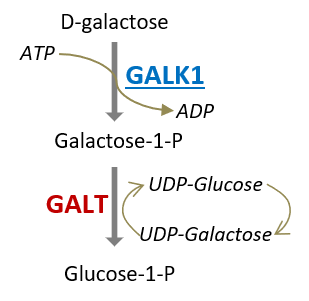
SMALL MOLECULE DRUG DISCOVERY
Disease-associated metabolic enzymes often lead to aberrant flux and accumulation of toxic metabolites. Therefore, pathway manipulation to reduce flux to the defective enzyme could have therapeutic benefit. Our objectives are to develop small molecule inhibitors for enzymes upstream of a metabolic defect, as ‘substrate reduction therapy’. Check out our hit discovery projects of GALK1, HAO1, ALAS2 and AASS in the public domain.
Northern Eye - A Cryo-Electron Microscopy Resource in the North East
2024
BBSRC Alert 23
Reconstruction and computational modelling for inherited metabolic diseases - Recon4IMD
2023-2027
Horizon Europe, UKRI Guarantee
Hit to lead development of GALK1 inhibitors for classic galactosemia
2024-2026
MRC Developmental Pathway Gap Fund
Developing small molecule inhibitors for a rare childhood seizure disorder
2021-2023
Action Medical Research and Life Arc
Hit-to-lead development of HAO1 inhibitors of primary hyperoxaluria
2021-2023
Harrington UK Rare Disease Scholar Award
Structural and biophysical characterisation of bicyclic peptides in the context with their therapeutic targets
2021-2023
Bicycle Therapeutics